- #Chromatogram viewer for mac how to
- #Chromatogram viewer for mac upgrade
- #Chromatogram viewer for mac full
- #Chromatogram viewer for mac code
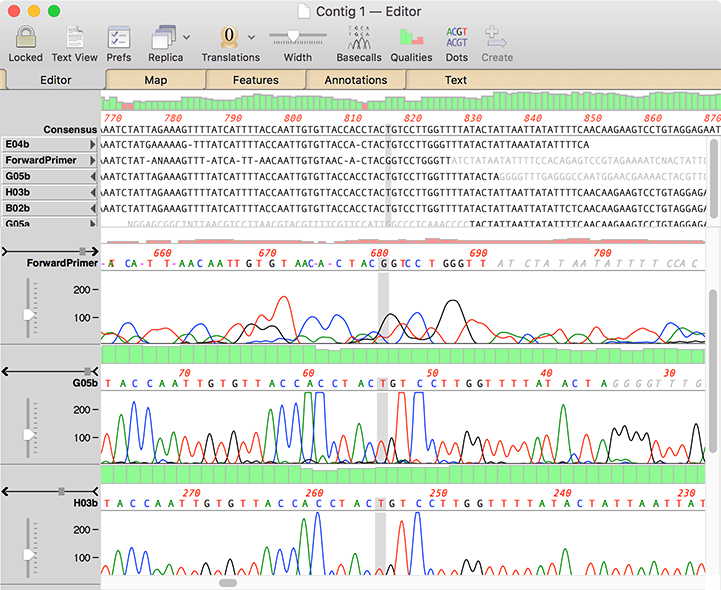
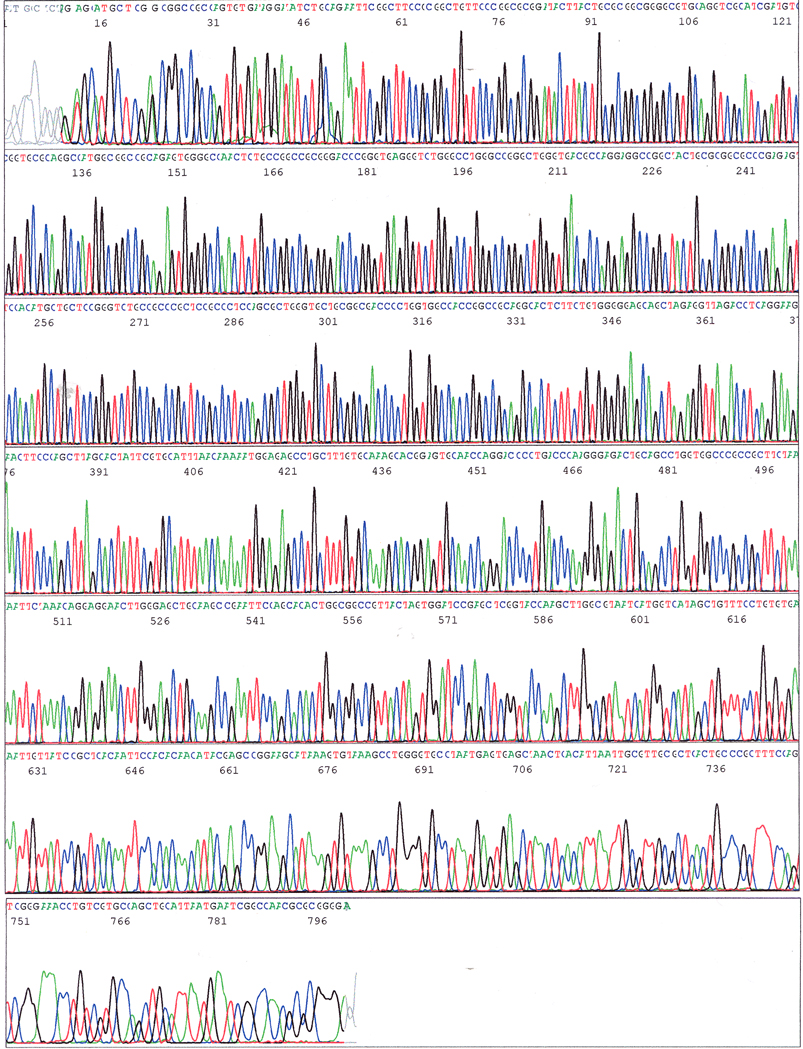
You should see individual, sharp and evenly spaced peaksģ. You can use any of the following programs to view your. Here are a few guidelines to help with DNA sequencing troubleshooting and analysis 1. In fact this is so ambiguous that the DNA sequencing reaction should be repeated. If you never looked at the trace you would be happy.īut look closer, the overlapping peaks in the chromatogram suggest the results are not as certain as the sequence may suggest. Here is an example of a seemingly clean DNA sequence (no Ns in sight). An example of where the chromatogram can come to your rescue for DNA sequencing troubleshooting and analysis And, like all controls, missing out is a big mistake. When it comes to DNA sequencing the chromatogram is your visual control. These controls help you properly visualize your results. When you run a restriction digest on a gel you always include proper controls like uncut DNA and the proper ladder. The most important of those is to always look closely at the trace file (or chromatogram) of the sequencing results you get back from your favorite sequencing facility. So I have developed some good habits that I wanted to pass on to you to make sure you are getting the most out of the data you get back from your sequencing runs.
#Chromatogram viewer for mac full
For full details, see the release history.As part of my job ensuring plasmid quality at Addgene, I analyze 50-100 sequencing reactions a week. SeqTrace 0.8.1 fixes several bugs present in the 0.8.0 release. You can read about all of the new features in SeqTrace 0.9.0 in the release history. I'd like to express my appreciation to all of the users who took the time to give me feedback on SeqTrace 0.8 your ideas and comments were very helpful in deciding what to focus on for 0.9.0.
#Chromatogram viewer for mac upgrade
This is a significant upgrade that includes many important improvements and new features as well as a few relatively minor bug fixes. It's been almost two years since the last release of SeqTrace, and after a lot of work, I'm very pleased to announce that SeqTrace 0.9.0 is now available. January 14, 2014: SeqTrace 0.9.0 released This will be the last release of SeqTrace to use the GTK+ 2 toolkit and PyGTK. For full details of the improvements in 0.9.1, see the release history. SeqTrace 0.9.1 also fixes a critical bug that caused some of the settings controls to display incorrectly under recent versions of the GTK+ toolkit on Linux. For many users, the most notable improvements are likely to be the various ways in which support for SCF input files has been enhanced. This new release does not introduce any major new features, but it does include several important improvements and bug fixes. SeqTrace 0.9.1 is now available for download. News January 11, 2018: SeqTrace 0.9.1 released
#Chromatogram viewer for mac how to
To get in touch with me directly, see the project information page to read more about SeqTrace and how to contact me.
To submit bug reports, please use SeqTrace's issue tracking system.
#Chromatogram viewer for mac code
I welcome feedback, bug reports, or code contributions from users. SeqTrace is written in Python, uses PyGTK with the GTK+ windowing toolkit, and runs on most popular operating systems, including GNU/Linux, Windows, and OSX. You can also see screen shots of SeqTrace running on other operating systems. To get started using SeqTrace, see the quick start guide, read the installation instructions, or browse the full user documentation.Ĭlick the image above to see a full scale screen shot of SeqTrace running on XUbuntu 11.10.
SeqTrace supports popular trace file formats, including ABIF, SCF, and ZTR. You can view your sequencing chromatograms at a variety of scales and zoom levels, simultaneously view matching forward and reverse traces, edit the called bases, and export individual DNA sequences as well as forward/reverse alignments. SeqTrace also includes a full-featured trace file viewer and editor. The finished DNA sequences can then be exported to common sequence file formats, such as FASTA. SeqTrace can automatically identify, align, and compute consensus sequences from matching forward and reverse traces, filter low-quality base calls, and perform end trimming of finished sequences. SeqTrace makes it easy to quickly generate high-quality finished sequences from a large number of trace files. SeqTrace is an application for viewing and processing DNA sequencing chromatograms (trace files).
